- NIH-supported, international study underscores importance of research of diverse populations
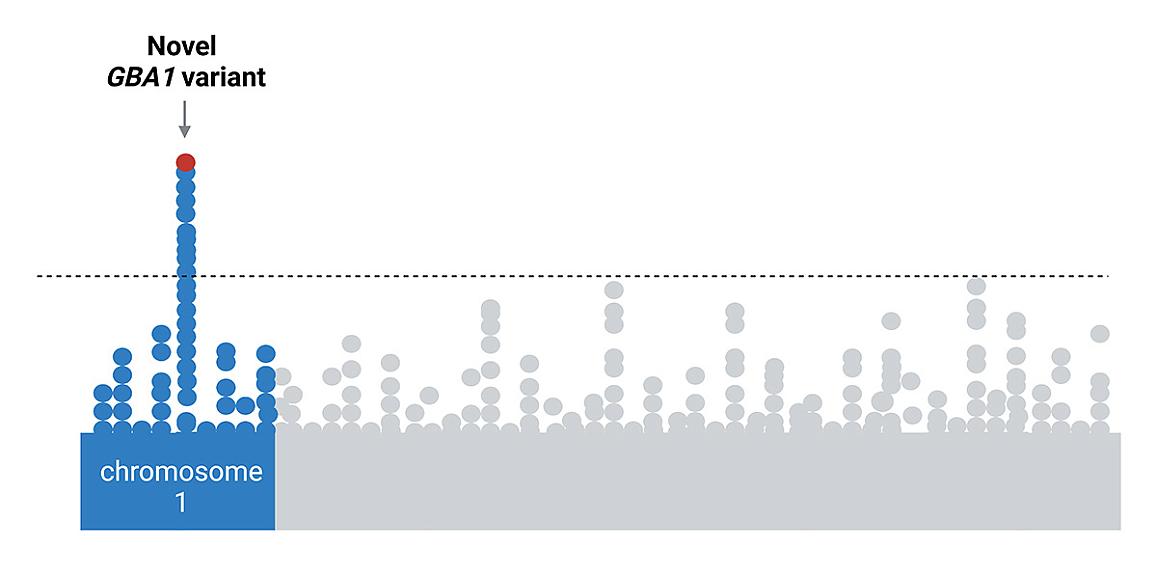
Maryland, USA: A gene variant found almost exclusively in the genomes of people of African ancestry increases the risk of developing Parkinson’s disease, according to an international study of nearly 198,000 participants with this genetic background. Published in The Lancet Neurology, the study results suggest the risk may be linked to a variant in the gene encoding β-glucocerebrosidase (GBA1), a protein known to control how cells in the body recycle proteins.
The study was led by scientists at the National Institutes of Health; the University College, London; and the University of Lagos, Nigeria. Although more research is needed to understand the role of environmental and other factors in these populations, the scientists found that those who carry one copy of the gene are about 1.5 times more likely to have Parkinson’s disease than those who have no copies whereas those who carry two copies are about 3.5 times more likely.
“To effectively treat Parkinson’s and truly any disease, we must study diverse populations to fully understand what the drivers and risk factors are for these disorders,” said Andrew B. Singleton, Ph.D., director, NIH Intramural Center for Alzheimer’s Related Dementias (CARD) and a study author. “These results support the idea that the genetic basis for a common disease can differ by ancestry, and understanding these differences may provide new insights into the biology of Parkinson’s disease.”
Over the past few decades, researchers have found several genetic risk factors for Parkinson’s disease. Rare inherited cases of the disease have been linked to about 20 genes harbouring pathogenic variants – formerly known as disease-causing mutations — while more than 100 regions of the human genome are associated with more common, sporadic forms of the disease. However, most of these findings are based on studies of people of European descent and very few have been conducted on people of African descent.
For this project, the researchers conducted a genome wide association study (GWAS) involving 1,488 people who had Parkinson’s disease and 196,430 people who did not. NIH scientists worked with researchers from around the world who are part of the Global Parkinson’s Genetics Program (GP2)(link is external), including the Black and African American Connections to Parkinson’s Study(link is external) and the International Parkinson Disease Genomics Consortium (IPDGC) – Africa(link is external). GP2 is supported by the Aligning Science Across Parkinson’s (ASAP)(link is external) initiative and implemented by The Michael J. Fox Foundation for Parkinson’s Research (MJFF)
Together the researchers collected DNA samples and analysed genetic data from individuals primarily from Nigeria and four sites across the United States. These data were combined with de-identified genetic and phenotypic data from 195,587 people of African American or Afro-Caribbean descent who consented to participate in 23andMe research.
A preliminary analysis of the data showed a significant association between Parkinson’s disease risk and the newly identified variant of the GBA1 gene. A review of previous studies also showed that this new variant rarely appears in people of European and Asian descent, suggesting it is almost exclusively linked to African ancestry.
“We were completely surprised. The goal of the initial analysis was to help train GP2 researchers in Nigeria and other parts of the world in how to analyse GWAS data,” said Sara Bandrés-Ciga, Ph.D., staff scientist at NIH CARD and an author of the study. “The fact that the GBA1 variant had a significant association while others did not suggest that it is strongly tied to Parkinson’s disease in this population.”
Further analysis of this study’s GWAS data suggested that the risk associated with the GBA1 variant is additive.
Previous studies by NIH researchers and others have shown that pathogenic variants of the GBA1 gene are also associated with Parkinson’s and Gaucher’s disease, a rare genetic disorder caused by problems with lysosomes. Lysosomes are tiny sacs inside of cells that break down proteins for recycling.
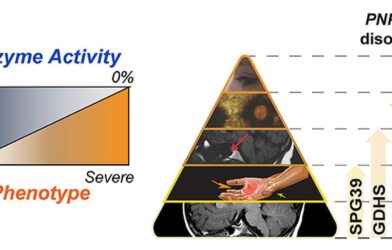
The results from this study suggest the new variant may alter lysosomal GBA1 activity in a previously unknown way. Most previously identified variants appear in the part of the genetic code that guides how the body manufactures glucocerebrosidase, an enzyme encoded by GBA1. This either disrupts the manufacturing process or alters the enzyme’s activity. In contrast, this newly discovered Parkinson’s disease variant appears just outside of the coding region. Further research is needed to determine how the variant may change activity.
Currently, researchers are developing genetic therapies for treating Gaucher’s disease and other lysosomal disorders.
“Our results represent a good first step towards fully understanding the genetic and biological complexity of each individual around the world who has Parkinson’s disease,” said Singleton. “Our hope is that results like these will provide researchers a roadmap for developing new genetic treatments and therapies for Parkinson’s disease.”
According to the World Health Organization(link is external), more than 8.5 million people around the world have Parkinson’s, a brain disorder that typically affects people 60 years of age or older. Initially, the disease can cause slowness, as well as balance and movement problems, including tremors and stiffness. Over time, the disorder may worsen, causing problems with walking, talking, sleeping, remembering, and mood.
Researchers can obtain data from the study by accessing the Accelerating Medicines Partnership® (AMP®)(link is external) program platform. The AMP® PD program is a public-private partnership managed by the Foundation for the National Institutes of Health and funded by the National Institute of Neurological Disorders and Stroke in partnership with the ASAP; Celgene Corporation, a subsidiary of Bristol-Myers Squibb Company; GlaxoSmithKline plc; MJFF; Pfizer Inc.; Sanofi US Services Inc.; and Verily Life Sciences.
–NIH news